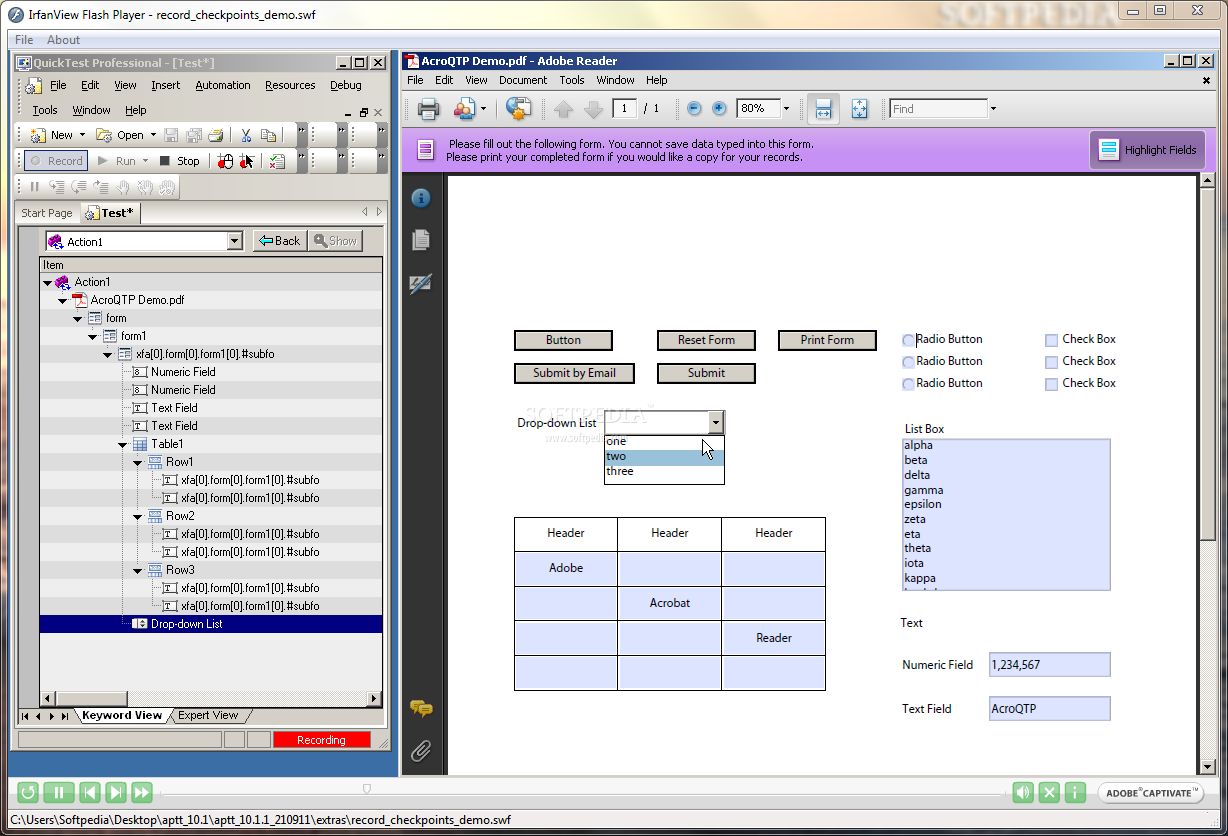
The programmable pre-crRNA, which contains nuclease guide sequences (spacers) interspaced by identical direct repeats, is processed to mature crRNA in combination with tracrRNA. The CRISPR/Cas system, which is employed in a variety of organisms, is derived from the Streptococcus pyogenes type II CRISPR system and consists of three genes, including one encoding Cas9 nuclease and two noncoding RNA genes: trans-activating crRNA (tracrRNA) and precursor crRNA (pre-crRNA). This system is based on the bacterial and archaeal clustered regularly interspaced short palindromic repeats (CRISPR) adaptive immune system for purging invading viral and plasmid DNA, which relies on the endonuclease activity of CRISPR-associated (Cas) proteins, with sequence specificity directed by CRISPR RNAs (crRNAs). Recently, another DSB-based breakthrough technology for genome editing, the CRISPR/Cas system, was developed. Based on DSBs at target loci, sequence-specific nucleases, including homing meganucleases, zinc finger nucleases, and transcription activator-like effector (TALE) nucleases have emerged as powerful technologies for targeted genome editing in eukaryotic organisms. DSBs can also result in homologous recombination (HR) between chromosomal DNA and foreign donor DNA through the HR pathway. Therefore, new technologies that are affordable, efficient, and user-friendly are needed for plant genome targeting.ĭouble-strand breaks (DSBs) at specific genomic sites can introduce a mutation at the DNA break site via the error-prone non-homologous end-joining (NHEJ) pathway. Moreover, T-DNA insertional mutants cannot be obtained for every gene of interest. However, the current method for generating plants carrying multiple mutated genes requires time-consuming and labor-intensive genetic crossing of single-mutant plants. There is an increasing demand for plants bearing mutations in multiple genes in order to dissect the functions of gene family members with redundant functions and to analyze epistatic relationships in genetic pathways. For the majority of researchers, transfer DNA (T-DNA) and transposon insertional mutagenesis remain the main sources of mutants of genes of interest in model plants such as the dicot Arabidopsis thaliana and the monocot rice ( Oryza sativa). We developed a toolkit that facilitates transient or stable expression of the CRISPR/Cas9 system in a variety of plant species, which will facilitate plant research, as it enables high efficiency generation of mutants bearing multiple gene mutations.Īpproaches for precise, efficient gene targeting or genome editing are highly important for functional genomic analysis of plants and for the production of genetically engineering crops. Moreover, the multiple-gene mutations could be inherited by the next generation. More importantly, using this toolkit, targeted mutations of three Arabidopsis genes were detected in transgenic seedlings of the T1 generation. The toolkit was validated using maize protoplasts, transgenic maize lines, and transgenic Arabidopsis lines and was shown to exhibit high efficiency and specificity. This toolkit requires no restriction enzymes besides BsaI to generate final constructs harboring maize-codon optimized Cas9 and one or more gRNAs with high efficiency in as little as one cloning step. We developed a CRISPR/Cas9 binary vector set based on the pGreen or pCAMBIA backbone, as well as a gRNA (guide RNA) module vector set, as a toolkit for multiplex genome editing in plants. Check whether a particular location on the page is empty, e.g.To accelerate the application of the CRISPR/Cas9 (clustered regularly interspaced short palindromic repeats/ CRISPR-associated protein 9) system to a variety of plant species, a toolkit with additional plant selectable markers, more gRNA modules, and easier methods for the assembly of one or more gRNA expression cassettes is required.splitting based on headings (requires PDFlib+PDI in addition to TET) Process PDFs based on their contents, e.g.  
Convert the contents of PDFs to other formats.Implement the PDF indexer for a search engine.TET contains advanced content analysis algorithms for determining word boundaries, grouping text into columns, identifying table structures and removing redundant items such as shadow text. TET optionally converts PDF documents to an XML-based format called TETML which contains text and metadata as well as resource information. Raster images are extracted in common image formats. TET makes available the text contents of a PDF as Unicode strings, plus detailed color, glyph and font information as well as the position on the page. PDFlib TET (Text and Image Extraction Toolkit) reliably extracts text, images, comments and metadata from PDF documents.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
